Research team will look at medical and social consequences

By Karyn Houston
Plant & Microbial Biology
Computational Biologist Steven Brenner will be part of an ambitious effort to assess whether large-scale gene sequencing aimed at detecting disorders and conditions can and should become a routine part of newborn testing.
Brenner, a professor in the Department of Plant & Microbial Biology at UC Berkeley, is part of a UC San Francisco team granted $6 million by the National Institutes of Health to identify the accuracy and feasibility of providing genetic sequencing as part of, or instead of, the current newborn screening that relies on biochemical changes in the blood. It also will assess what additional information would be useful to have at birth and the ethics and public interest in having such tests performed.
"Genome sequencing has the potential to reduce costs and improve the effectiveness of newborn screening," Brenner said, allowing early intervention for infants and fundamental changes in the tehcnology for screening newborns.
Goals
The project has three broad goals:

- Find out if DNA sequencing works as well or better than current tests used for newborn screening
- Find out whether the low marginal cost of screening for additional illnesses with DNA sequencing allows this technology to improve on what is currently being done. "These are diseases for which we expect there would be agreement that it would be appropriate to screen for, but there is no present test that is affordable," Brenner said.
- Research the ethical, legal, and social consequences of whole genome sequencing in newborn screening, and the additional sorts of information it could reveal, such as how well a child will respond to different drugs. "Is this something that would be valuable for parents and clinicians to know, and would it be reasonable and appropriate to consider expanding the remit of newborn screening this way?" Brenner said.
Enormous Implications
The cost of genome sequencing has plummeted but the ethical and moral questions surrounding genetic testing loom large.
The first genome sequenced, about a decade ago, cost nearly $3 billion. Today scientists can sequence all of an individual's genes for a few hundred dollars. The Brenner Lab, experts at computational biology, are at the forefront of this new area of science.
"This dramatic change is having an enormous impact throughout biological science and biomedical research, and is beginning to have profound impacts" in the medical community, Brenner said.
Newborn screening has been a given in hospitals for more than 50 years, to recognize babies with serious diseases that might not be noticed in the newborn period but where early awareness and intervention can have a major impact. Genome sequencing has the potential to reduce costs, expand testing and improve outcomes.
“Genomic sequencing has the potential to diagnose a vast array of disorders and conditions at the very start of life,” said Alan E. Guttmacher, MD, director of the Eunice Kennedy Shriver National Institute of Child Health and Human Development (ICHD), which is jointly funding the studies. “But the ability to decipher an individual’s genetic code rapidly also brings with it a host of clinical and ethical issues, which is why it is important that this program explores the trio of technical, clinical and ethical aspects of genomics research in the newborn period.”
Precision Medicine
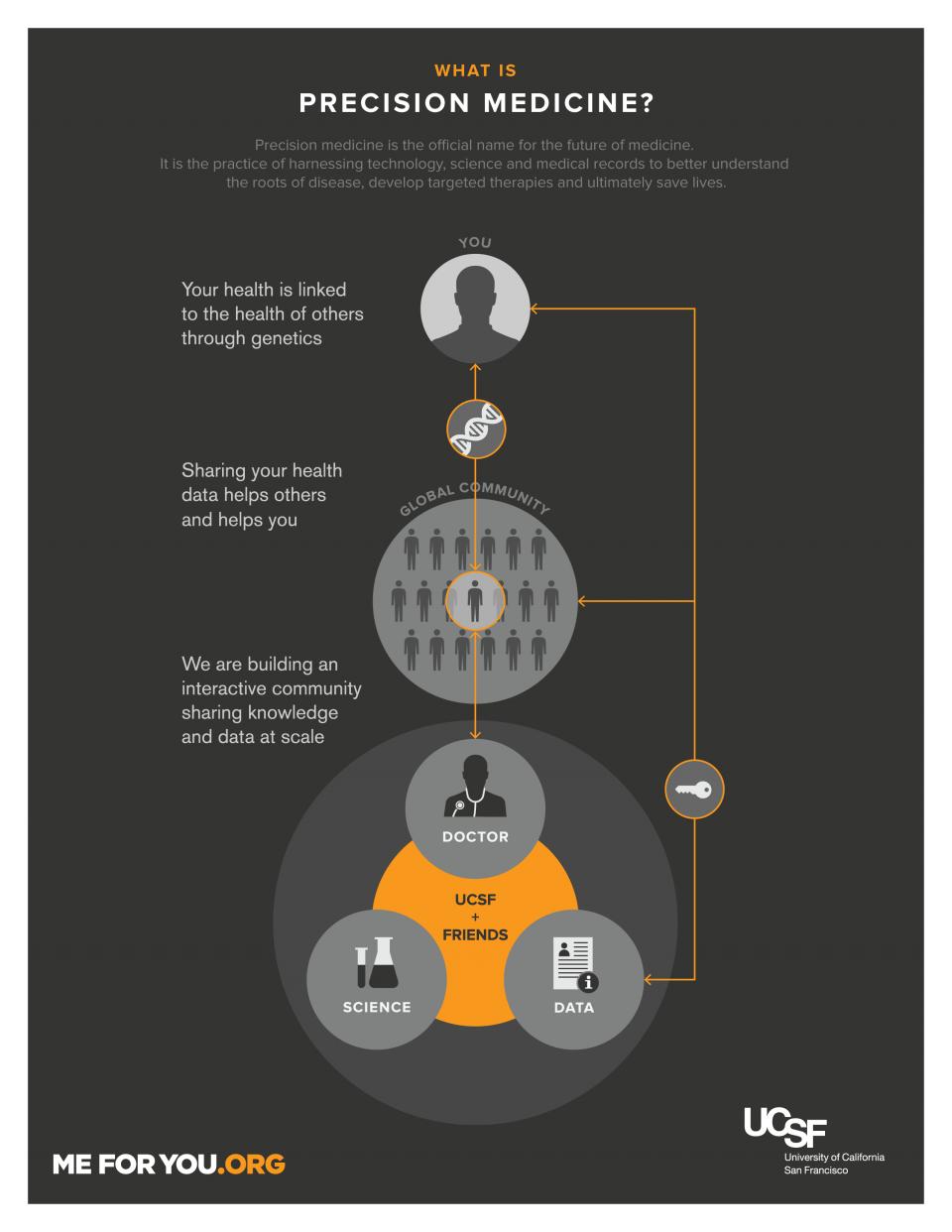 The research is a core element in the emerging field of precision medicine, which aims to harness vast amounts of genetic and health data to create predictive, preventive and precise care for patients on an international scale. Doing so has the potential to transform medicine, but there are many logistical and ethical hurdles to resolve along the way.
The research is a core element in the emerging field of precision medicine, which aims to harness vast amounts of genetic and health data to create predictive, preventive and precise care for patients on an international scale. Doing so has the potential to transform medicine, but there are many logistical and ethical hurdles to resolve along the way.
The UCSF team will study the potential of sequencing the exome – the roughly 2 percent of DNA that represents genes which code for proteins – as a method of newborn screening. The research will look at the exome’s potential for identifying disorders that California currently includes in the newborn screen, as well as those that are not currently screened for, but for which newborns may benefit if detection can occur early in life.
Brenner Team
Brenner's bioinformatics role will be in analyzing the genome sequences.
"We will be involved in first taking the raw data from the sequencers and processing it to obtain high quality information about the genetic variation in the newborns being screened," Brenner said. In collaboration with others the lab will then research whether this technology can be used to identify children with serious childhood diseases currently screened for, or related to those currently screened for.
The most commonly used DNA sequencers today provide short fragments of sequence information. Brenner's role in the project will be to first use a variety of techniques and software tools to determine, from short fragments, the nearly complete sequence of the genes. Interpreting this variation remains a difficult prospect. As part of another research effort, the Brenner Lab has led an international experiment (called the Critical Assessment of Genome Information) to determine the state-of-the art in the field.
"We will use databases of knowledge about what genetic variations lead to disease alongside computational methods to determine likelihood of the screened diseases from a genome sequence," Brenner said. "We will then evaluate whether these inferences lead to the same conclusions as present newborn screening approaches and clincial assessments."
Group Approach

The team will look at the issues from three vantage points:
The first is a partnership with the California Department of Public Health to test blood drops previously collected from 1,400 children statewide who received standard newborn screening, to determine whether exome sequencing would be more accurate and also whether it woud provide insights that could lead to improved newborn screening, care and treatment.
“My hope is that this will give us solid information on the specificity of gene testing, versus standard biochemical testing, for the disorders we are already screening for,” said Robert Nussbaum, MD, who leads the UCSF Division of Medical Genetics and holds the Holly Smith Distinguished Professorship in Science and Medicine at UCSF.
The second project will offer genetic testing to patients in a UCSF immune system disorders clinic run by Jennifer Puck, MD, a pediatrician in the UCSF Benioff Children’s Hospital whose research laboratory pioneered the current newborn test for Severe Combined Immunodeficiency (SCID). Parents will be asked to give informed consent for this arm of the project. Brenner and Puck share another recently-awarded NIH grant to use exome sequencing to understand the genetic basis of disease in newborns found from current newborn screens to have perious undiagnosed immunological disease. And the Brenner group has already used exome analysis in conjunction with Puck to successfully inform the diagnosis of newborns.
While there are several known genetic mutations that lead to the immune disorder, Puck’s original test simply looks at a marker of whether children lack the immune cells known as T lymphocytes, which are missing in SCID.
This new project will enable the team to assess whether exome sequencing (Brenner's area of expertise) works as well or better than the current test in identifying SCID, as well as other immune system abnormalities that the current test does not cover. Exome sequencing may also give parents information on the genetic basis of their child’s disease.
The third part of the project will offer parents genetic testing for newborns at the UCSF Benioff Children’s Hospital to assess whether the child is likely to have adverse reactions to medications based on their genetics – an area known as pharmacogenomics. That portion will be conducted in conjunction with UCSF ethicist Barbara Koenig, PhD, who will be studying parent’s attitudes regarding testing children beyond what is currently offered in newborn screening.
While the first two projects are mainly looking at whether genetic testing would be more accurate, specific and useful than current methods, this third element assesses how willing parents are to get genetic information about their child that may be useful later in life, but not right away.
“So far, newborn screening programs have not been directed towards just letting people know about a possible disease risk. There has to be a high probability of serious illness that can be prevented with early intervention,” Nussbaum said. “Pharmacogenomics is perhaps the most acceptable of tests that imply potential risk. There’s very little risk, and the possibility of great benefit, to knowing whether you will react to a drug or an anesthetic, and the only way to find out besides genetic screening is if you’re in the operating room or have filled a prescription and you have a bad reaction.”
Exploring Legal Issues
 The research team also intends to develop a participant protection framework for conducting genomic sequencing during infancy and will explore legal issues related to using genome analysis in newborn screening programs. Together, these studies have the potential to provide public health benefit for newborns and research-based information for policy makers.
The research team also intends to develop a participant protection framework for conducting genomic sequencing during infancy and will explore legal issues related to using genome analysis in newborn screening programs. Together, these studies have the potential to provide public health benefit for newborns and research-based information for policy makers.Additional researchers on the project include Neil Risch, PhD, director of the UCSF Institute of Human Genetics; Pui-Yan Kwok, MD, PhD, a UCSF professor of dermatology whose research focuses on analysis of complex genetic traits; and Joseph Shieh, MD, PhD, an assistant professor of pediatrics and medical genetics. Sean Mooney, PhD, a bioinformatics expert at the Buck Institute for Research on Aging, and Steven Brenner, PhD, a professor of plant and microbial biology at UC Berkeley and an adjunct professor at UCSF, will contribute their expertise in bioinformatics to the project.
The four NIH pilots, which also include Brigham and Women’s Hospital in Boston, Children’s Mercy Hospital in Kansas City, and the University of North Carolina at Chapel Hill, will receive $25 million over the next five years as funds are made available through the NICHD and the National Human Genome Research Institute, both parts of the National Institutes of Health. This year’s grants were made under the Genomic Sequencing and Newborn Screening Disorders research program.
The grant is part of a larger NIH initiative, the Genomic Sequencing and Newborn Screening Disorders research program, to test whether genetic sequencing is superior to current practices of newborn screening, as well as whether it is ethical. A total of $25 million will fund the initiative over the five-year period.
Important Links
San Francisco Business Times article about the research
Critical Assessment of Genome Interpretation websiite